
Robert P. Gunsalus, Ph.D.
1602 Molecular Science Building
Microbial Cell Metabolism in Bacteria and Archaea
Affiliations:
Professor Emeritus and Distinguished Research Professor,
Department of Microbiology, Immunology & Molecular Genetics
Faculty Member:
The UCLA DOE Institute
The UCLA Molecular Biology Institute
UCLA Graduate Programs in Biosciences (GPB) Home Areas:
Immunity, Microbes & Molecular Pathogenesis GPB
Genetics & Genomics GPB
Biochemistry, Biophysics & Structural Biology GPB
Research Interests :
Our current research focuses on the molecular biology and physiology of anaerobic microorganisms including the syntrophic bacteria, their associated methanogenic partners plus other archaeal species. These microbes colonize a wide range of habitats on Earth and possess the ability to adapt and grow over a wide range of environmental conditions (e.g., substrate type, temperature, pH, salinity). We are examining how their metabolism, energy harvesting and associated cell envelope structures adapt to these changes using a variety of genetic, molecular and biochemical approaches.
1) Genomics and Proteomics of Syntrophic Bacteria. The syntrophic bacteria along with their methanogenic archaeal partners are expert in the anaerobic recycling of biomass in marine and fresh water environments. However, we know relatively little about their metabolism, cell structures or their ability to adapt to changing conditions. Using genomic, transcriptomic and proteomic tools we are elucidating core metabolic pathways for carbon utilization and energy harvesting in model syntrophic microorganisms plus their methanogenic community partners. The longer-term goal is to generate predictive metabolic models that describe these co-operative partnerships between the syntrophic bacteria and methanogenic archaea.
Metabolic reconstruction of Syntrophomonas wolfei
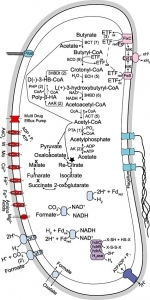
2) Structure and Function of Microbial Cell Envelopes. We are studying the composition, structure and function of microbial cell envelopes of representative archaeal and bacterial species. Cell envelopes may contain a protein coat or surface layer that surrounds the cell membrane that aids in protecting it from environmental challenges. We have isolated and characterized the cell envelopes of representative archaeal methanogenic strains of the genus Methanosarcina. The major envelope proteins were identified and where many were previously annotated to be “hypothetical proteins”. The most abundant envelope proteins in three of these strains, M. mazei, M.acetivorans and M. barkeri, were structurally characterized. Following cytoplasmic synthesis, each protein is post-translationally modified and then exported to the cell surface where they are assembled into an outermost cell envelope layer. This coat surrounds and protects the cytoplasmic membrane. The first high resolution three-dimensional structure of one of these archaeal S-layer proteins was recently obtained. These structure and function studies along with associated cell appendages (flagella and pili) provide insight regarding the ability of these archaea cell types to adapt to diverse growth conditions.
Methanospirillium hungatei cells and flagellum
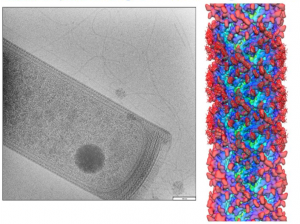
Biography:
Dr. Gunsalus is currently a Distinguished Research Professor in the Department of Microbiology, Immunology, and Molecular Genetics at the University of California, Los Angeles within the Schools of Life Sciences and Medicine. He obtained B.S. degrees is in Microbiology and in Chemistry from the South Dakota State University and went on to do his PhD studies in Microbial Biochemistry at the University of Illinois at Urbana Champaign working with Dr. Ralph S. Wolfe. Prior to joining the faculty at UCLA, he did postdoctoral work at Stanford University in Palo Alto in the areas of Molecular Biology with Dr. Charles Yanofsky and Dr. Robert D. Simoni.
Publications:
James, K.L., Kung, J.W., Crable, R.R., Mouttaki, M., Sieber, J.R., Nguyen, H.H., Yang, Y., Xie, Y., Erde, J., Wofford, N.Q., Karr, E.A., Loo, J.A., Ogorzalek Loo, R.R., Gunsalus, R.P. and M.J. McInerney. 2019. Syntrophus aciditrophicus uses the same enzymes in a reversible manner to degrade and synthesize aromatic and alicyclic acids. Environmental Microbiology. 21:1833-1846.
Lauren E. Cook, L.E., S. Gang, A. Ihlan, M. Maune, R.S. Tanner, M.J. McInerney, G. Weinstock, E.A. Lobos and R.P. Gunsalus. 2018. Genome sequence of Acetomicrobium hydrogeniformans OS1. Genome Announc. 2018 Jun; 6(26): e00581-18. Published online 2018 Jun 28. doi: 10.1128/genomeA.00581-18
Briege, A., C.M. Oikonomou, Y-W Chang, Kjaer,A., A.N. Huang, K.W. Kim, D. Ghosal, D. Kenney, R.O.O. Loo, R.P. Gunsalus, and Grant J. Jensen. 2017. “Morphology of the archaellar motor and associated cytoplasmic cone in Thermococcus kodakaraensis. EMBO Rep. Sep;18(9):1660-1670.
Poweleit, N., P. Ge, H.H. Nguyen, R.R. Ogorzalek Loo, R. Gunsalus, and Z.H. Zhou. CryoEM structure of the Methanospirillum hungatei archaellum reveals structural features distinct from the bacterial flagellum and type IV pilus. Nature Microbiology. 2016. 2: 16264.
Gunsalus, R.P., L.E. Cook, B. Crable, L. Rohlin, E. McDonald, H. Mouttaki, J.R. Sieber, N. Poweleit, Z.H. Zhou, A.L. Lapidus H.E. Daligault, M. Land, P. Gilna, N. Ivanova, N. Kyrpides, D.E. Culley and M.J. McInerney. Complete genome sequence of Methanospirillum hungatei type strain JF1. Standards in Genomic Sciences. 2016; 11: 2.
Toso D.B, M.J. Javed, E. Czornyj, R.P. Gunsalus, and Z.H. Zhou. Discovery and Characterization of Iron Sulfide and Polyphosphate Bodies Coexisting in Archaeoglobus fulgidus Cells. Archaea. 2016. 2016: 4706532.
Arbing, M.A., S. Chan, A. Shin, T. Phan, C.J. Ahn, L. Rohlin, and R.P. Gunsalus. Structure of the surface layer of the methanogenic archaean Methanosarcina acetivorans. Proceedings of the National Academy of Sciences of the United States of America. 2012; 109: 11812-7.
Sieber, J.R. M.J. McInerney and R.P. Gunsalus. Genomic Insights into Syntrophy: The Paradigm for Anaerobic Metabolic Cooperation. Annual Review of Microbiology. 2012. 109(29): 429-452.
McInerney, M.J. L. Rohlin, H. Mouttaki, U.M. Kim. R.S. Krupp, L. Rios-Hernandez, J.R. Sieber, C.G. Struchtemeyer, A. Bhattacharyya, J.W. Campbell, and R.P. Gunsalus. The genome of Syntrophus aciditrophicus: life at the thermodynamic limit of microbial growth. Proceedings of the National Academy of Sciences of the United States of America. 2007; 104: 7600-5.
Lab Members:
Shichun Ma, – Visiting Professor
Erin Natalie – Research Assistant
Shirley Zhang – Undergraduate Researcher
Derek Lam – Undergraduate Researcher
Megan Chu– Undergraduate Researcher
Links:
UCLA Molecular Biology Institute
UCLA DOE Institute of Genomics and Proteomics
UCLA Medical School Biosciences
http://bioscience.ucla.edu/faculty/robert-p-gunsalus
EcoCyc – the Escherichia coli protein gene database
Departmental Graduate Studies in Microbiology
http://bioscience.ucla.edu/admissions-overview
Contact:
Dr. Robert Gunsalus
Department of Microbiology, Immunology, and Molecular Genetics
UCLA
609 Charles E Young Dr S
1602 Molecular Science Building
Los Angeles, CA 90095
(310) 206-8201 office
robg@microbio.ucla.edu
